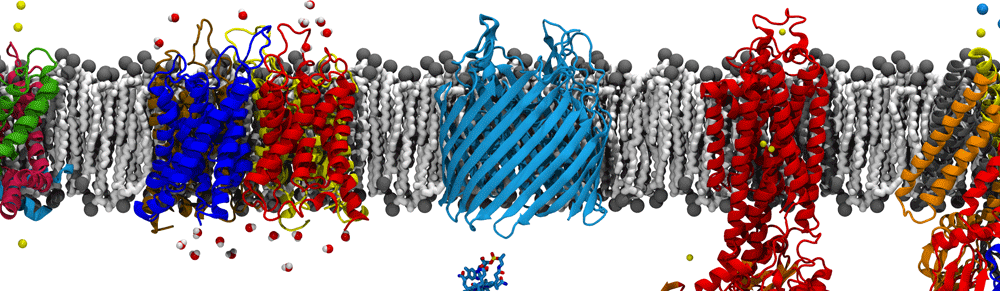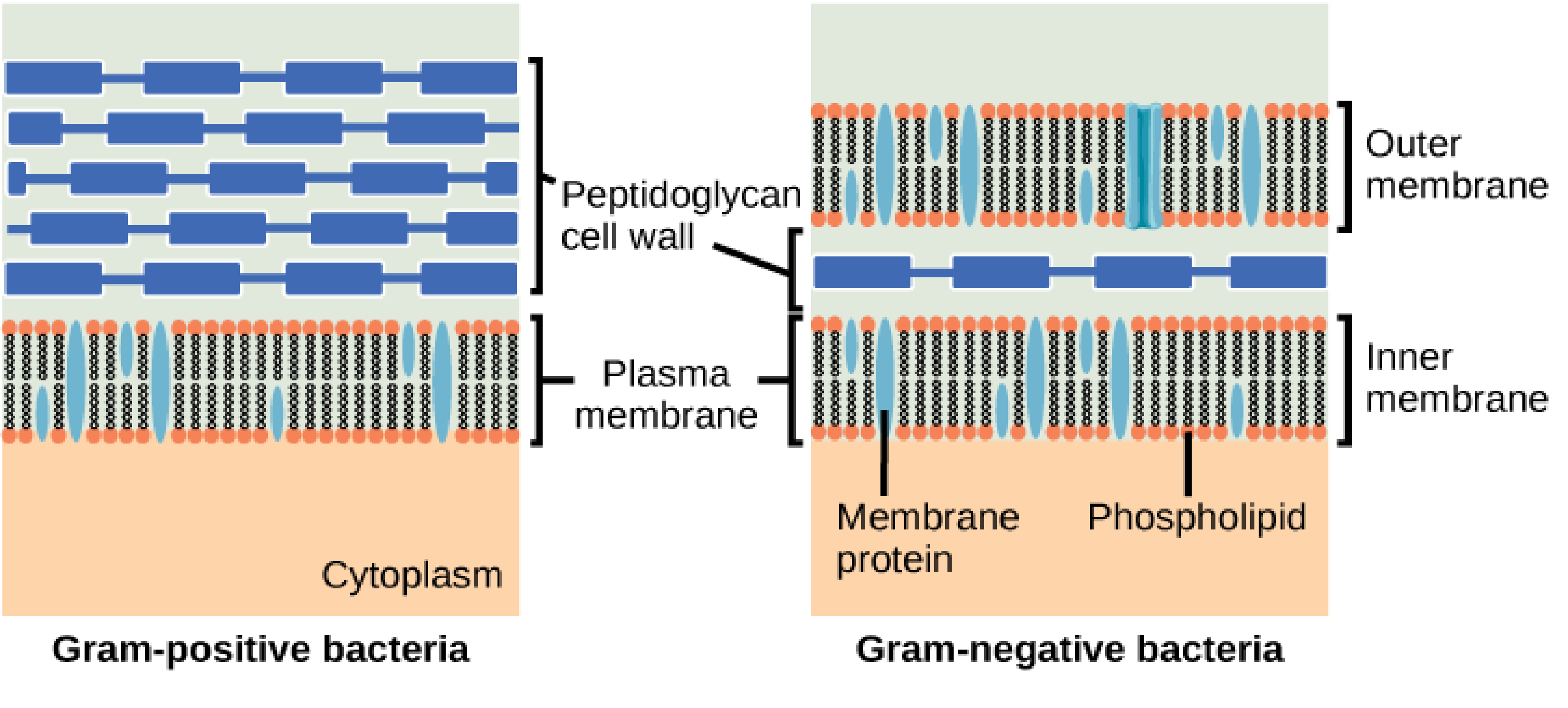
Schematic differences between Gram-positive (left) and Gram-negative (right) bacteria. The former have a single bilayer with an enveloping peptidoglycan cell wall protecting the bacterium, whereas the latter have two cell membranes separated by a periplasm region, within which is the cell wall.
The outer membrane of Gram-negative bacteria (those possessing two enveloping membranes with a periplasm in between, as seen above right) is unique for a number of a reasons. First and foremost is the outer membrane composition. While it possesses the typical two-tailed phospholipids in its inner leaflet, the outer leaflet is composed of lipopolysaccharides (LPS). LPS molecules have 5-6 acyl tails combined with a chain of sugar molecules stretching out above the membrane (see the figure below). Certain species of Gram-negative bacteria can also contain O-antigen chains protruding from the core oligosaccharide.
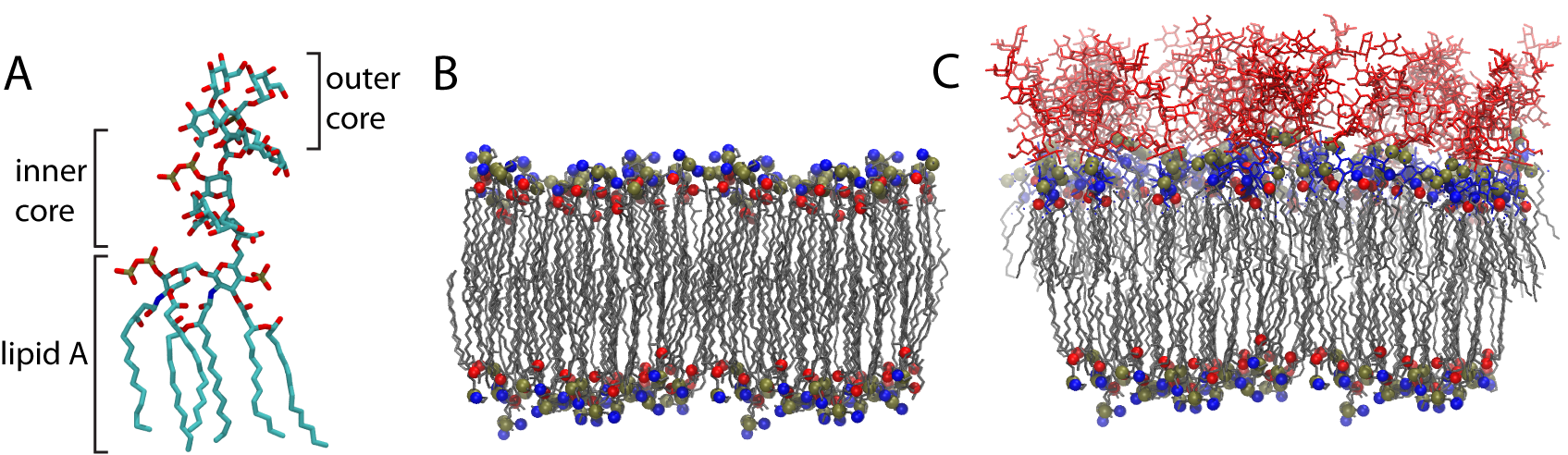
(A) an example LPS molecule. The inner & outer core composition varies within and between species (B) a standard, amphiphilic, phospholipid bilayer. (C) a model outer membrane with LPS in the outer (top) and phospholipids within the inner (bottom) leaflets .
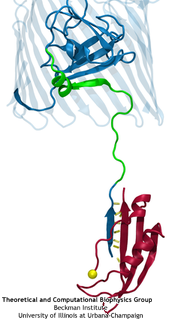
TonB (red) “pulling” on the lumenal domain of the transporter BtuB (blue), the connection being mediated by hydrogen bonds (yellow).
Another unique aspect of the outer membrane is that it’s full of porins, which permit the diffusion of small molecules relatively easily across it. As such, energy cannot be generated or stored here. Nonetheless, bacteria often need to actively import large and/or scarce nutrients across the outer membrane. To get energy for this task, they utilize an inner membrane protein, TonB, that couples across the periplasm to an outer membrane transporter for import of, e.g., vitamin B12 or iron extracted from human transferrin. The mechanisms of this interaction and how it causes transport are still open questions in the field of microbiology. We have proposed that energy is transduced through a mechanical coupling between the inner and outer membrane complexes, supported by simulations that demonstrate the strength of the TonB-transporter connection mediated by hydrogen bonds, as well as the response of the transporter to the application of force. In particular, we see in multiple contexts that the interior domain of the transporter unfolds, generating a pathway for the substrate across the membrane.
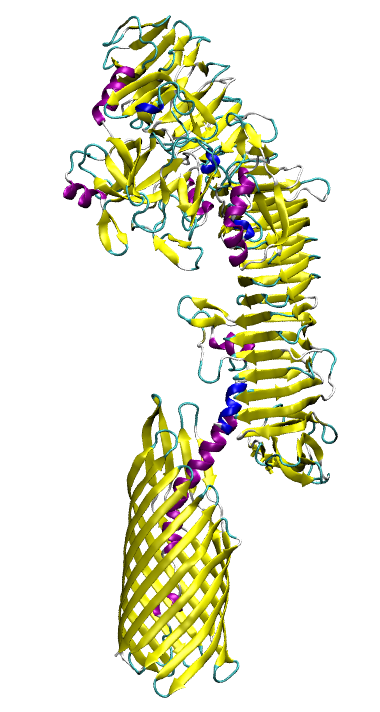
Just as bacteria need to import molecules across the outer membrane, they sometimes need to export them as well. For example, certain virulence factors are secreted through the use of so-called “auto-transporters”, proteins that export themselves! The auto-transporter consists of two domains, a membrane-bound barrel at the C-terminus and a secreted domain at the N-terminus. After transport, the N-terminal domain is typically self-cleaved, freeing it from the membrane. We have carried out simulations on the native pre-cleaved state as well as on a number of biochemically characterized mutants. The simulations reveal an unusual conformation of the catalytic asparagine residue, seen in less than 2% of all atomic structures, that is supported by the unique electrostatic environment created by neighboring residues. This conformation is predicted to greatly accelerate the cleavage reaction, and the other residues found to be involved are in agreement with experimental observations.
Publications
- Structural basis for iron piracy by pathogenic Neisseria.
N. Noinaj, N. C. Easley, M. Oke, N. Mizuno, J. Gumbart, E. Boura, A. N. Steere, O. Zak, P. Aisen, E. Tajkhorshid, R. W. Evans, A. R. Gorringe, A. B. Mason, A. C. Steven, and S. K. Buchanan. Nature, 483:53-58, 2012. - Molecular basis for activation of a catalytic asparagine residue in a self-cleaving bacterial autotransporter.
T. J. Barnard, James Gumbart, J. H. Peterson, N. Noinaj, N. C. Easley, N. Dautin, A. J. Kuszak, E. Tajkhorshid, H. D. Bernstein, and S. K. Buchanan. Journal of Molecular Biology, 415:128-142, 2012. - Coupling of calcium and substrate binding through loop alignment in the outer membrane transporter BtuB.
J. Gumbart, M. C. Wiener, and E. Tajkhorshid. Journal of Molecular Biology, 393:1129-1142, 2009. - Mechanics of force propagation in TonB-dependent outer membrane transport.
J. Gumbart, M. C. Wiener, and E. Tajkhorshid. Biophysical Journal, 93:496-504, 2007.
Featured in Science Magazine’s Editors’ Choice: Science, 316:1101, 2007.