Education
- PhD in Computational Chemistry, University of Amsterdam, Netherlands (2013)
- MSc in Chemical Engineering, Royal Institute of Technology, Stockholm, Sweden (2008)
Research
Stalling of the bacterial ribosome by macrolides
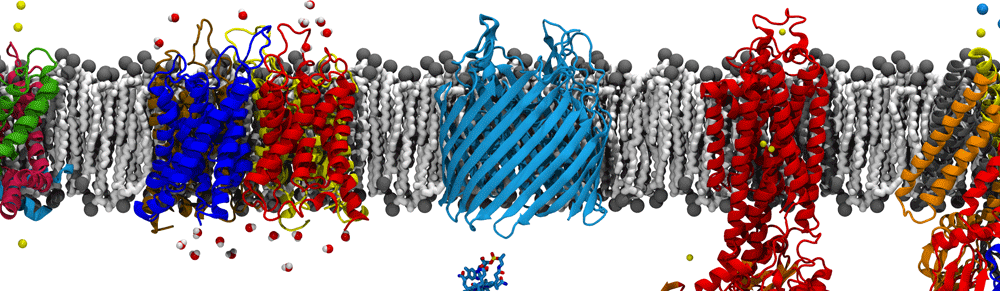
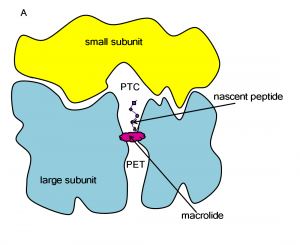
The ribosome is a large molecular machine, consisting of rRNA and proteins, that is responsible for protein synthesis in all cells. During protein synthesis, the genetic information in mRNA is decoded into a sequence of amino acids and these amino acids are linked together with peptide bonds, one at a time. Both the recognition and linking of amino acids occur in the peptidyl transferase center (PTC), while the nascent peptide exits the ribosome through the protein exit tunnel (PET). Since there are significant structural differences between eukaryotic and prokaryotic ribosomes, the ribosome makes an excellent target for antibiotics, which can kill harmful bacteria without harming the host eukaryotic cells.
One class of antibiotics that targets the bacterial ribosome is macrolides and macrolide derivatives. Their general structure is a 14-, 15- or 16-atom cyclic lactone ring with attached sugars, peptides or other functional groups .
Macrolides operate by binding to the ribosome in the PET and block the exit of the nascent peptide. The blockage prevents continued synthesis once the nascent peptide reaches the macrolide and the unfinished peptide dissociates from the ribosome. Unfortunately, due to the extensive use of macrolides, several strains of bacteria have become resistant to them. According to experimental studies, exchange or methylation of specific rRNA residues in the ribosome induces resistance to some or all macrolides.
We use molecular simulations to understand how specific rRNA mutations result in macrolide resistance and to facilitate the development of compunds that can overcome this resistance.
Previous Publications
- Clarifying the role of sodium in the silica oligomerization reaction. Anna Pavlova, Thuat T. Trinh, Rutger A. van Santen and Evert Jan Meijer Phys. Chem. Chem. Phys. 15, 1123-29 (2013).
- Understanding the Role of Water in Aqueous Ruthenium-Catalyzed Transfer Hydrogenation of Ketones. Anna Pavlova and Evert Jan Meijer ChemPhysChem 13(15), 3492-96 (2012)
e-mail: anna.pavlova// at // physics.gatech.edu office: W207 Howey Physics Building mail: School of Physics, 837 State Street, Atlanta, GA 30332-0430